Inside Angle
From 3M Health Information Systems
Making time for change: The importance of updating terminologies
Laboratorians are trained to use exact measures in quality to provide the right answer on the right patient at the right time. This attention to precision is the cornerstone in supporting a wide variety of clients who are dedicated to using the LOINC and SNOMED CT terminologies to report laboratory test results. When recent interactions with the public at a LOINC meeting revealed that some labs may not continue to take a proactive stance after adopting the standard terminologies required to achieve Meaningful Use incentives, the HDD lab team (consisting of four ASCP-certified clinical laboratory scientists) decided to take a look over the fence and see what might be at stake. If downstream health information exchange processes were programmed to operate off of LOINC terms, what happens to unrecognized codes being sent from an updated system to a non-updated system? This is an important data flow process to understand because nightmares form from unresolved error logs that prevented information from posting on a patient record.
In order to investigate the effect of outdated terminology versions, the HDD lab team performed a retrospective analysis of terminology changes across LOINC and SNOMED CT versions. The retrospective included a comparison of biannual releases over the past four years for LOINC and SNOMED CT. Eight releases for each standard were reviewed: SNOMED CT (July 2012 – January 2016) and LOINC versions 2.40 to 2.54 (June 2012 – December 2015). The team used differential views between each release to calculate the actual content increase per department for LOINC. The microbiology specialist also relied on her client reports of new SNOMED CT terms for existing phrases in microbiology reports. Organism interaction within laboratory reports were main focus in studying SNOMED CT releases (although SNOMED CT can be used for specimens and sources documentation). Since a picture holds a thousand words, the following figures speak for the variety of new terms in LOINC since version 2.40 (released in June 2012).
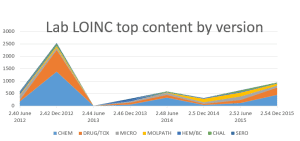
The positive slope starting at June 2013 shows continued release of new content after the 2.42 release in December 2012, with almost a 60 percent increase over the 2.42 release coming between 2012 and 2015. There’s no question that more LOINC terms are generated. The original question of the retrospective was “What’s the harm of not updating a system interacting with other systems that are updating with each release?” By resorting this same content, but changing from a date focus to a department/category focus, the team was given a perspective of types of terms added to the lab LOINC database. This leads to Figure 2.
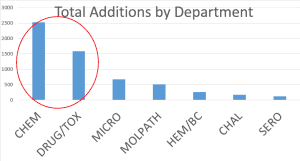
The largest amount of new content (over 4,000 terms) is recognized in chemistry and drug/toxicology. Chemistry and medications can literally impact organ function, system function and human life in a heartbeat. These lab values may have greatest impact on immediate, critical patient care. The HDD lab team concurred with the lab LOINC committee meeting at Regenstrief: Healthcare systems risk miscommunication if they do not update from the original mandates of LOINC 2.40 or SCT July 2012. If codes are unrecognized by an outdated system, the lab result message may go into an error log, instead of posting directly to a patient’s record. Having an incomplete patient record is a patient safety issue.
The HDD Lab team also experienced clients questioning validity of SNOMED CT codes provided them. The team uses all of SNOMED CT, and upon investigation with the client, found the local laboratory information system (LIS) was constraining SNOMED CT to a few domains (a microbiology module of the LIS might only use Organisms, Qualifiers and Specimens domains, for example). By leaving the Findings domain out of LIS usability, many common terms were excluded. Specifically, negative isolate observations and organism differentiation are being placed in the Findings domain, rather than Organism domain.
Further, more than 200 negative isolation observations exist as of January 2016 release.A “ruling out” option is frequently used in obtaining the correct diagnosis and subsequent treatment plan. Healthcare systems risk miscommunication if processes or systems segregate the SCT database by domains, causing restrictions in code searching. The microbiology result message of “Group A Streptococcus not isolated” (SCT ID 704664007) may go into error log and not post. Conversely, terms added to the Findings domain may not get the message out that “Clostridium difficile toxin A detected” (SCT ID 310541005) in an LIS system that constrains microbiology reports. Clostridium infection risk factors in the hospital and long term care facilities list exposure in close settings. What if it’s unknown the adjoining location has an infectious diarrhea? Again, missing information due to a system’s outdated release platform is a patient safety issue.
It’s clear to see that vocabulary standards are very dynamic and under constant development. Thus, it is crucial to complete annual audits as part of your maintenance plan. You may find new LOINC terms better suited, realizing the correct intent of vocabulary coding.
Here are a few highlights the HDD lab team has found pertinent to all of our clients. Did your site catch and update these already?
- LOINC is creating new methods for the BioFire arrays with a melting curve method; they should not be mapped to probe.amp.tar. (expected June 2016)
- LOINC discouraged methodless LDH codes, as there is a 3-fold difference in values between lactate to pyruvate vs pyruvate to lactate substrates.
- LOINC deprecated methodless d-dimer codes as there is 2-fold DVT decision thresholds between Fibrinogen Equivalent Units (FEU) and D-Dimer Units (DDU).
If local resources at your site are strung too tight, and performance is stalled, feel free to contact 3M with questions or direction in providing the right codes to the client’s system at the right time.
Pamela Banning, BS, is a senior healthcare data analyst for 3M Health Information Systems.


